 Image credit: Authors
Image credit: AuthorsAbstract
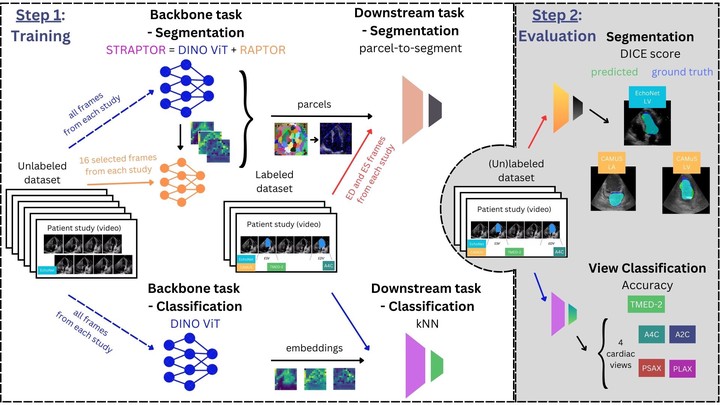
Supervised training of deep learning models for medical imaging applications requires a significant amount of labeled data. This is posing a challenge as the images are required to be annotated by medical professionals. To address this limitation, we introduce the Adaptive Locked Agnostic Network (ALAN), a concept involving self-supervised visual feature extraction using a large backbone model to produce anatomically robust semantic self-segmentation. In the ALAN methodology, this self-supervised training occurs only once on a large and diverse dataset. Due to the intuitive interpretability of the segmentation, downstream models tailored for specific tasks can be easily designed using white-box models with few parameters. This, in turn, opens up the possibility of communicating the inner workings of a model with domain experts and introducing prior knowledge into it. It also means that the downstream models become less data-hungry compared to fully supervised approaches. These characteristics make ALAN particularly well-suited for resource-scarce scenarios, such as costly clinical trials and rare diseases. In this paper, we apply the ALAN approach to three publicly available echocardiography datasets, EchoNet-Dynamic, CAMUS, and TMED-2. Our findings demonstrate that the self-supervised backbone model robustly identifies anatomical subregions of the heart in an apical four-chamber view. Building upon this, we design two downstream models, one for segmenting a target anatomical region, and a second for echocardiogram view classification.